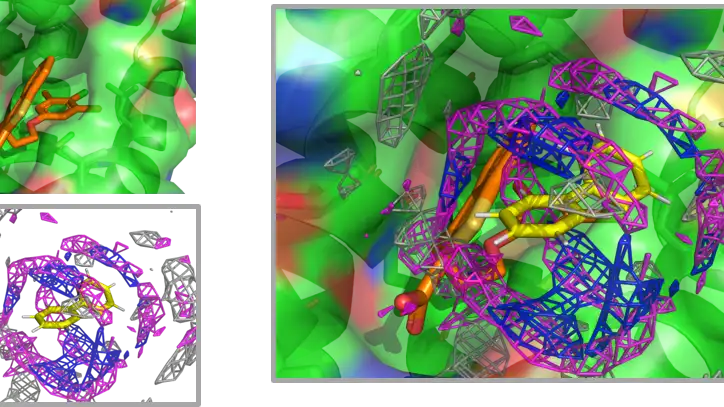
AAp-MSMD
Amino Acid Preference Mapping using Mixed-Solvent MD
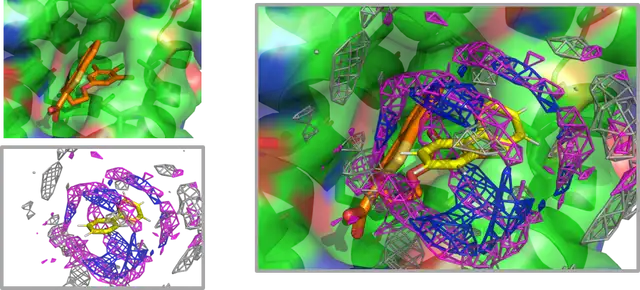
AAp-MSMD is a method that uses mixed-solvent molecular dynamics (MSMD) simulations with various amino acids as probes to predict peptide-binding sites on proteins and estimate changes in binding affinity caused by single-residue substitutions in peptides.
This approach can be applied to estimating protein-protein interactions (PPI) and protein-peptide binding, and is expected to contribute to new drug discovery modalities.